Quantitative Multiplexed ChIP Reveals Global Alterations that Shape Promoter Bivalency in Ground State Embryonic Stem Cells - ScienceDirect

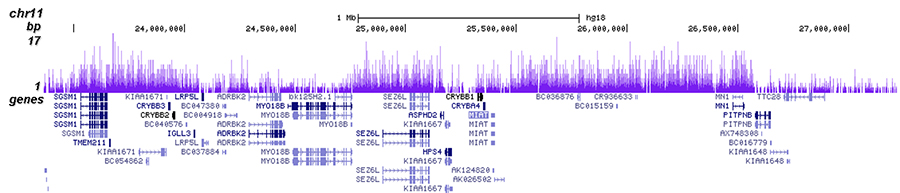
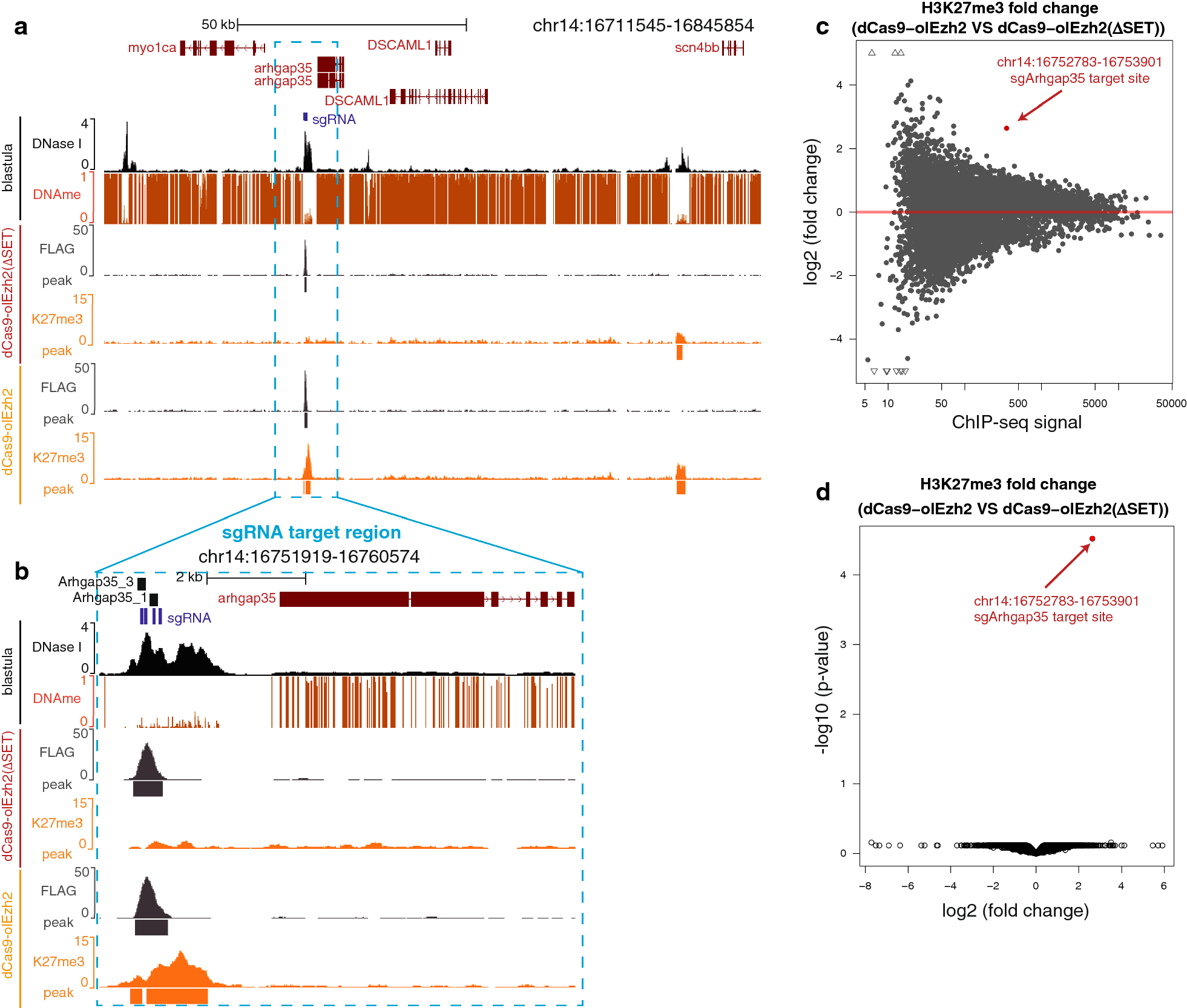
H3K27me3-rich genomic regions can function as silencers to repress gene expression via chromatin interactions | bioRxiv

Local difference of H3K27me3 marks in mutants of PcG components. (A)... | Download Scientific Diagram

Global changes of H3K27me3 domains and Polycomb group protein distribution in the absence of recruiters Spps or Pho | PNAS

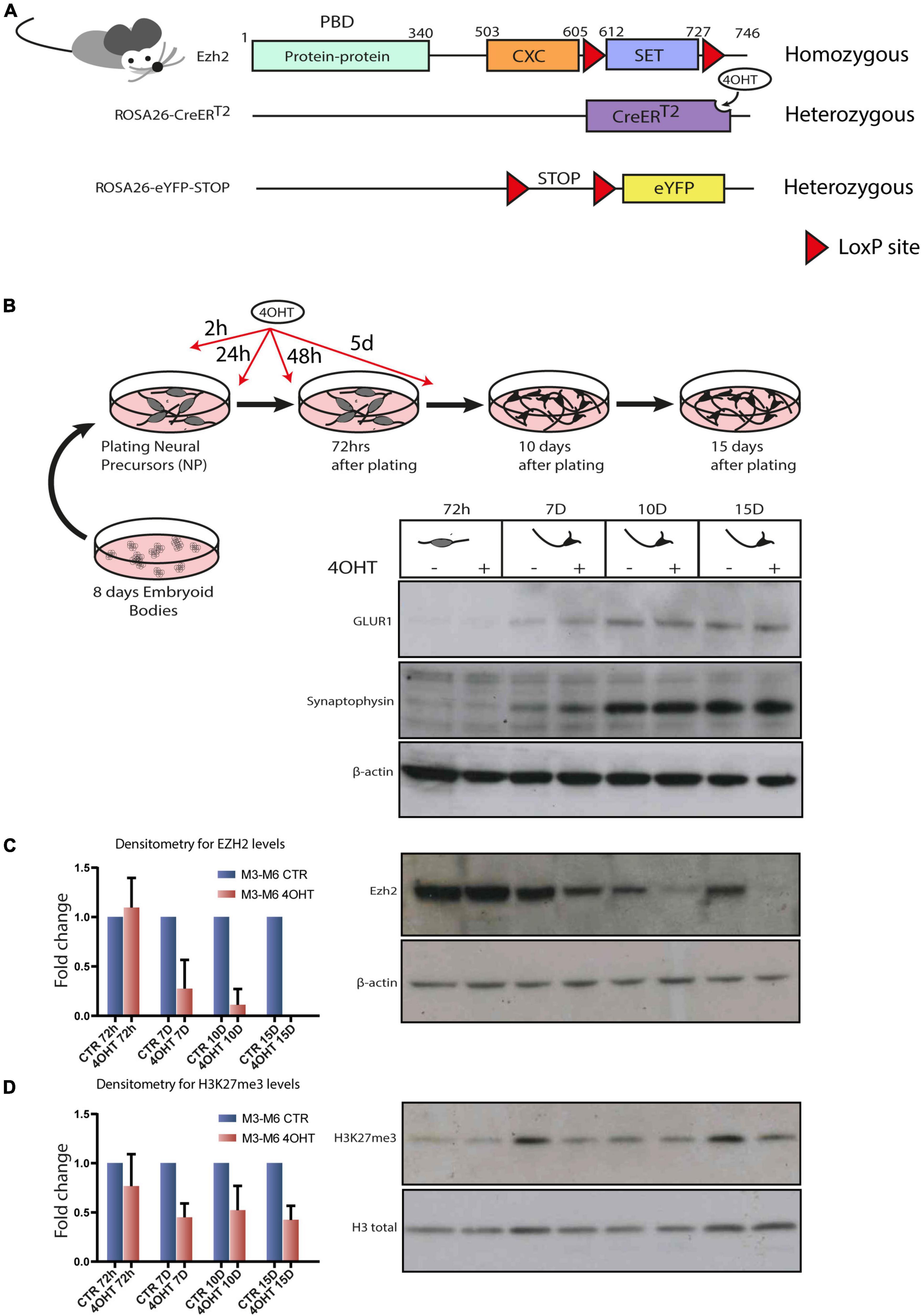
Epigenomics-Based Identification of Major Cell Identity Regulators within Heterogeneous Cell Populations - ScienceDirect

Genome‐wide occupancy of histone H3K27 methyltransferases CURLY LEAF and SWINGER in Arabidopsis seedlings - Shu - 2019 - Plant Direct - Wiley Online Library
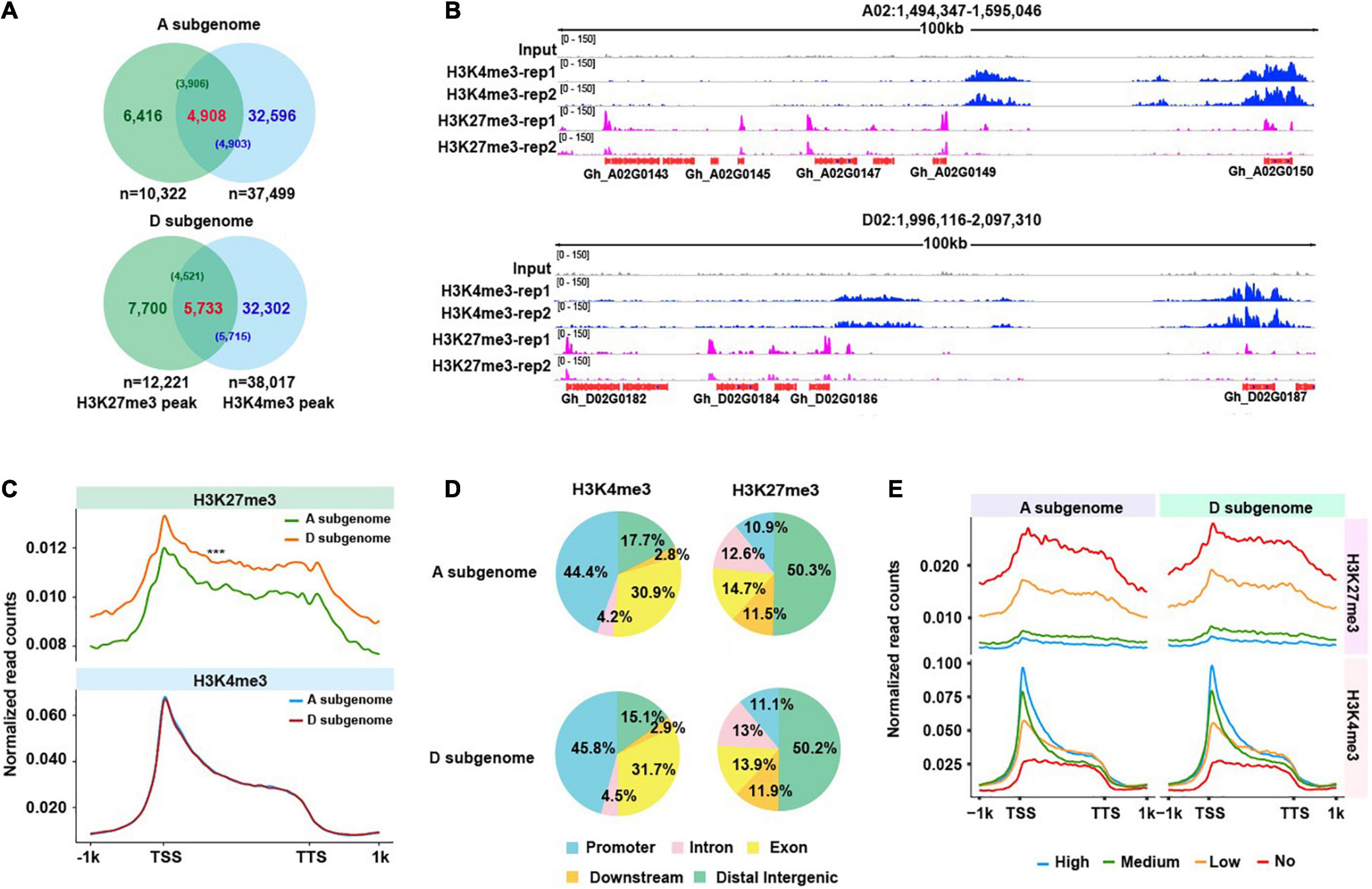
Frontiers | Profiling of H3K4me3 and H3K27me3 and Their Roles in Gene Subfunctionalization in Allotetraploid Cotton
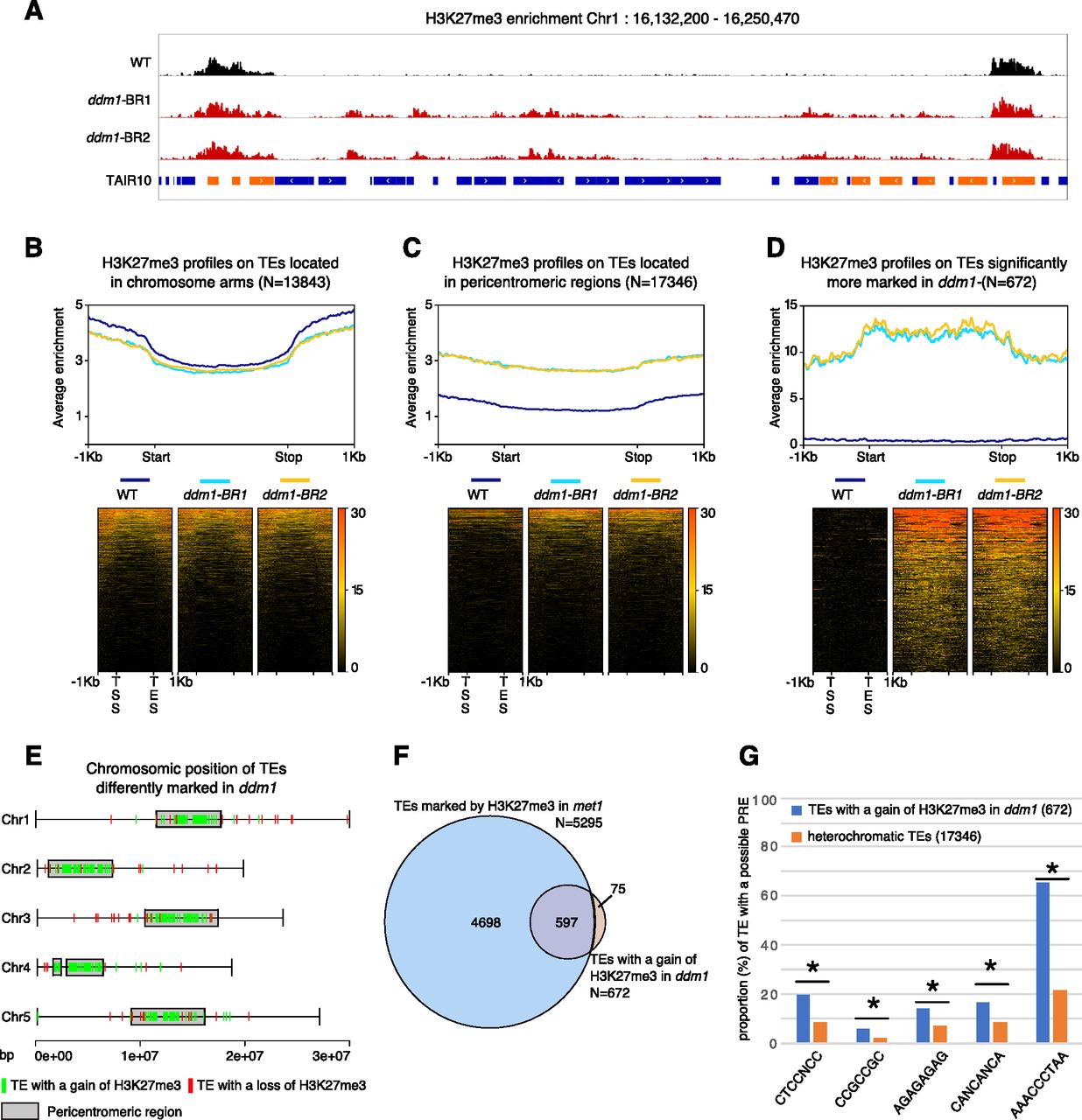
Polycomb mutant partially suppresses DNA hypomethylation–associated phenotypes in Arabidopsis | Life Science Alliance
Genomewide Analysis of PRC1 and PRC2 Occupancy Identifies Two Classes of Bivalent Domains | PLOS Genetics
Genomewide Analysis of PRC1 and PRC2 Occupancy Identifies Two Classes of Bivalent Domains | PLOS Genetics

Figure 2 from ChIP-seq analysis reveals distinct H3K27me3 profiles that correlate with transcriptional activity | Semantic Scholar

ChIP-seq of EZH2 and H3K27me3 in WT iPSCs a, Comparison of the EZH2... | Download Scientific Diagram
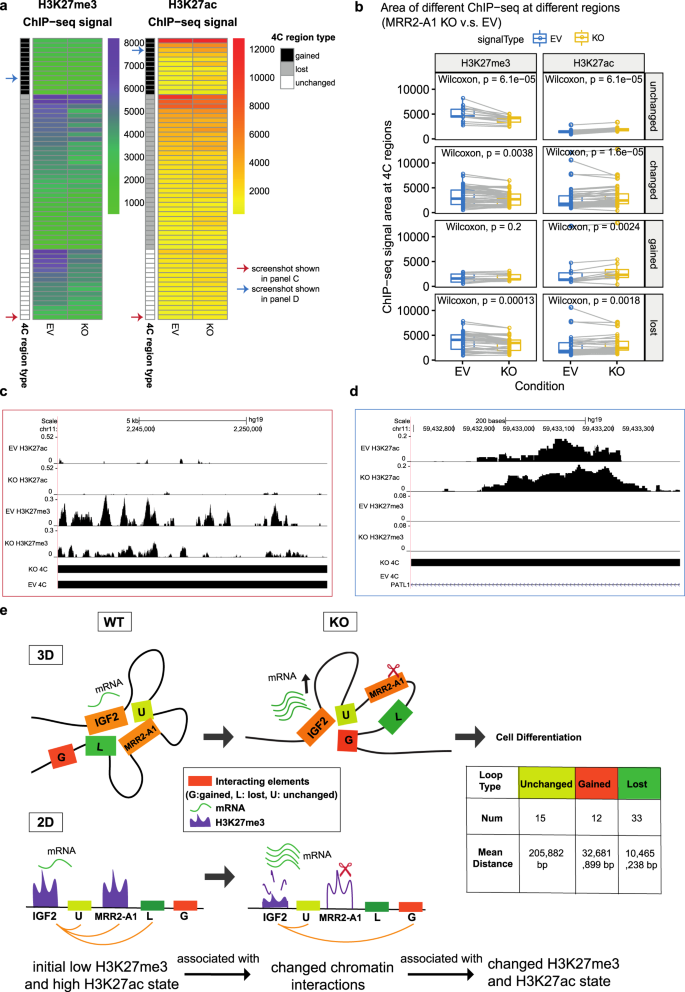
H3K27me3-rich genomic regions can function as silencers to repress gene expression via chromatin interactions | Nature Communications

Genome-wide localization patterns by ChIP-seq of H3K27me3, Su(z)12, and... | Download Scientific Diagram

H3K27me3-rich genomic regions can function as silencers to repress gene expression via chromatin interactions | bioRxiv
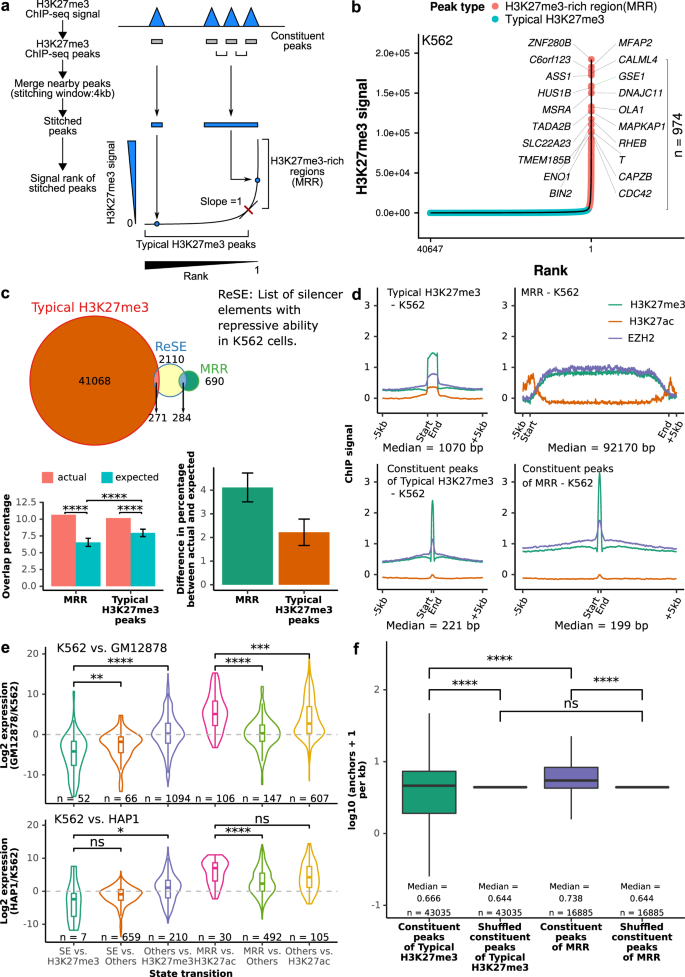
H3K27me3-rich genomic regions can function as silencers to repress gene expression via chromatin interactions | Nature Communications

ChIP-seq analysis to test chromatin occupancy of FIE, H3K27me3, AZF1... | Download Scientific Diagram